Doctoral position
Equivariant transformers for 3D molecular reconstruction in cryo-EM
Deadline: March 8, 2024
Contact: janden@kth.se
Cryogenic electron microscopy (cryo-EM) is a powerful imaging technique for reconstructing 3D models of biological macromolecules using transmission electron microscopy. Although this method is able to reach near-atomic resolution for certain molecules many important challenges remain. Foremost is the issue of noise. Due to the low electron dose required to reduce specimen damage, cryo-EM images are extremely noisy, a fact that is exacerbated for small molecules that have a low signal power to begin with. To compensate for this, a large number of images (often in the hundreds of thousands) are required to obtain an accurate 3D reconstruction of the molecule. However, these large datasets can be quite expensive to obtain and for small enough molecules the noise is too high that reconstruction is impossible using current methods.
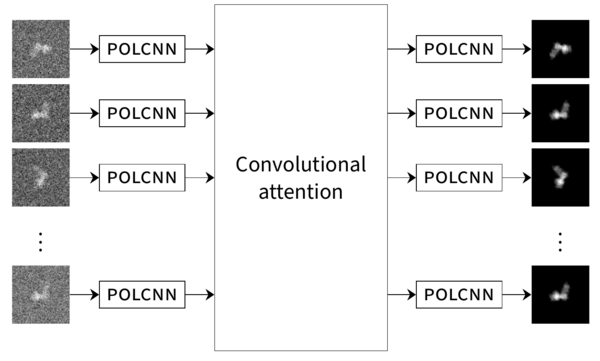
This project instead proposes to mitigate the high noise level by training deep neural networks (DNNs) to encode important the relevant structure in the data (cryo-EM images and reconstructed molecular models). Due to the geometry of the problem, these networks need to satisfy certain rotational and translational equivariance contraints since projection images can be translated and rotated arbitrarily without affecting the underlying molecular structure. In addition, the redundant nature of cryo-EM datasets naturally lends itself to transformer architectures, which have the potential to recombine information from the various images in order to reduce the noise level (denoising) and ultimately create accurate 3D molecular models (reconstruction).
The goal of this project is therefore to first construct an equivariant transformer architecture that can be trained on experimental cryo-EM images to provide superior denoising accuracy. The next step is to attach this transformer to another equivariant DNN based on a method-of-moments estimator to yield a 3D reconstruction of the molecule. The resulting method will allow for more accurate reconstructions at high noise levels, reducing the expense of data collection and enabling reconstruction of small molecules. Finally, the method will also enable accurate and robust reconstruction of structural variability from moderate-sized datasets, something which is not possible using current methods.
The successful doctoral candidate will be part of a larger team of researchers working on different aspects of deep learning and cryo-EM imaging at KTH Department of Mathematics. Beyond this core team, the student will also have the opportunity to work with collaborators elsewhere in Sweden and internationally. The student will be enrolled in the Doctoral program in Applied and Computational Mathematics specializing in mathematical statistics and will also be involved in teaching duties for various courses in the department. This position is a full-time, five-year position starting in August 2024 or at some other date to be agreed upon.
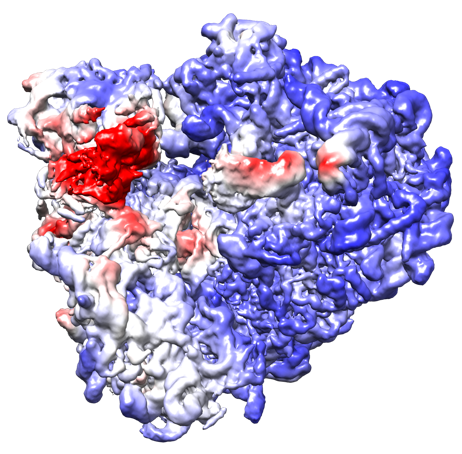
D. Kimanius et al. Exploiting prior knowledge about biological macromolecules in cryo-EM structure determination. IUCrJ 8:60–75, 2021.
A. Vaswani et al. Attention is all you need. Proc. NeurIPS, 2017.
J. Andén and A. Singer. Structural varaibility from noisy tomographic projections. SIAM Journal on Imaging Science, 11(2):1441–1492, 2018.
N. Sharon, J. Kileel, Y. Khoo, B. Landa, and A. Singer. Method of moments for 3D single particle ab initio modeling with non-uniform distribution of viewing angles. Inverse Problems 36(4):04403, 2020.
J. Zhao and A. Singer. Rotationally invariant image representation for viewing direction classification in cryo-EM. Journal of Structural Biology, 186(1):153–166.